Decomposition of Ligand-Binding Affinity
时间:2022-09-05
Xiaohui Wang, Bin Chong, Zhaoxi Sun, Hao Ruan, Yingguang Yang, Pengbo Song, and Zhirong Liu*.
More is simpler: decomposition of ligand-binding affinity for proteins being disordered.
Protein Sci. 31 (7), e4375 (2022).
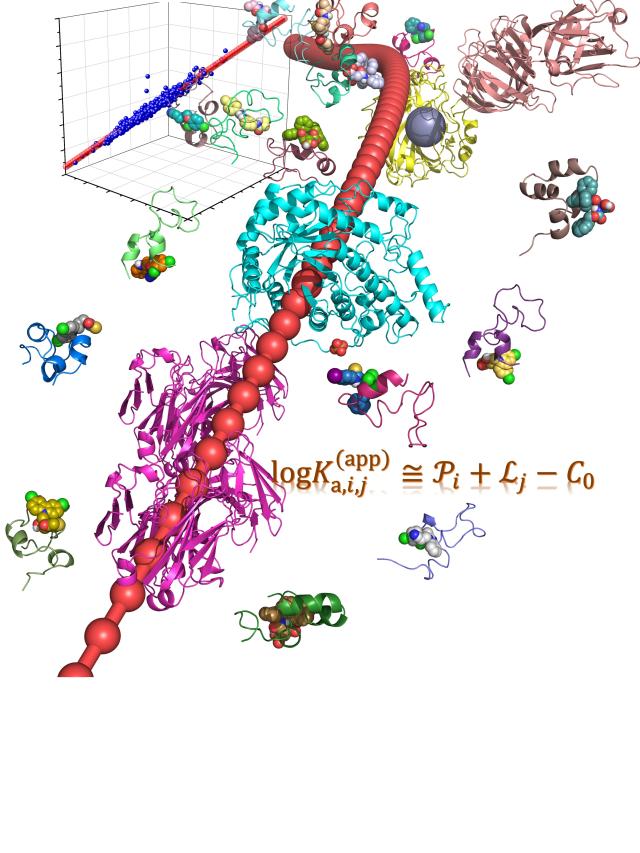
Abstract: In statistical mechanics, it is well known that the huge number of coupled degrees of freedom does not complicate the problem as it seems, but actually greatly simplifies the analysis (e.g., to give a Boltzmann distribution). Here, we reveal that the ensemble averaging from the vast conformations of intrinsically disordered proteins (IDPs) greatly simplifies the nature of binding affinity, which can be reliably decomposed into a sum of the ligandability of IDP and the capacity of ligand. Such an unexpected regularity is applied to facilitate the virtual screening upon IDPs. It also provides essential insight in understanding the specificity difference between IDPs and conventional ordered proteins since the specificity is caused by deviation from the baseline behaviour of protein-ligand binding.
Statement for a broader audience: The molecular world is full of noise and fluctuation, and the law of large numbers conquers the randomness to give simple results such as Boltzmann distribution. Here it is revealed that the ensemble averaging from the vast conformations of intrinsically disordered proteins greatly simplifies the nature of ligand binding affinity.
Link: https://onlinelibrary.wiley.com/doi/full/10.1002/pro.4375




