Reinforcement learning to boost molecular docking
时间:2021-01-15
Bin Chong, Yingguang Yang, Zi-Le Wang, Han Xing, and Zhirong Liu*.
Reinforcement learning to boost molecular docking upon protein conformational ensemble.
Phys. Chem. Chem. Phys. 23 (11), 6800-6806 (2021).
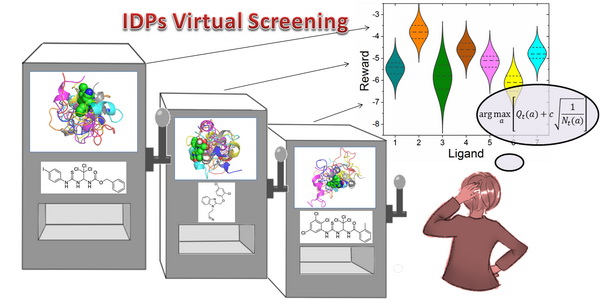
Abstract: Intrinsically disordered proteins (IDPs) are widely involved in human diseases and thus are attractive therapeutic targets. In practice, however, it is computationally prohibitive to dock large ligand libraries to thousands and tens of thousands of conformations. Here, we propose a reversible upper confidence bound (UCB) algorithm for the virtual screening of IDPs to address the influence of the conformation ensemble. The docking process is dynamically arranged so that attempts are focused near the boundary to separate top ligands from the bulk accurately. It is demonstrated in the example of transcription factor c-Myc that the average docking number per ligand can be greatly reduced while the performance ismerely slightly affected. This study suggests that reinforcement learning is highly efficient in solving the bottleneck of virtual screening due to the conformation ensemble in the rational drug design of IDPs..
Link: https://pubs.rsc.org/en/content/articlelanding/2021/cp/d0cp06378a#!divAbstract
亮点工作介绍:AI助力无序蛋白药物虚拟筛选,多臂老虎机再显神通




