The teams led by Prof. Chu Wang at the College of Chemistry and Molecular Engineering, Peking University, and Prof. Xiao-Dong Su at State Key Laboratory of Protein and Plant Gene Research, and Biomedical Pioneering Innovation Center (BIOPIC), Peking University have developed a coevolution-based computational pipeline named “MetalNet” to systematically predict metal-binding sites in proteomes. It provides a unique and enabling tool for interrogating the hidden metalloproteome and studying metal biology. The research results were published online by the journal Nature Chemical Biology on January 2, 2023 (https://www.nature.com/articles/s41589-022-01223-z), with the title "Co-evolution-based prediction of metal-binding sites in proteomes by machine learning ".
Proteins bind to metal ligands to undertake important physiological functions in living organisms, including structural stabilization, catalysis of biochemical reactions, storage of essential elements, transcriptional regulation, signal transduction, etc. Discovery and functional characterization of novel metal-binding proteins in proteomes will therefore be of great interest for both basic biology and industrial applications in the post-genome era. However, discovering novel metalloproteins by computational methods without sequence or structural homology from proteomes remains challenging.
Recently, the coevolution signals between residues calculated from multiple sequence alignments have been fed to machine learning models to guide accurate modeling of protein structures and protein-protein interactions. Inspired by such an approach, the authors have explored the distribution of coevolution signals in metal-binding sites and developed a computational method to predict metal-binding sites based on machine learning using coevolution signals.
The authors took the frequency matrix of residue type for every residue pair as input, and used metal-binding CHED coevolved pairs (as positive samples) and non-metal-binding CHED coevolved pairs (as negative samples) to train machine learning models. The final model works well, with an average AUC of 0.88 and an F1-score of 0.66. Considering that metal-binding sites often involve multiple residues in coordination, the authors assembled the residue pairs predicted by the machine learning model into a residue coevolution network, and obtained a relatively complete network cluster through a graph theory-based filter, which greatly improves the confidence of the prediction results and increases the F1-score to 0.72. Overall, the method requires only MSA-derived coevolutionary information as input and does not rely on any sequence or structural motifs to make predictions (Fig. 1).
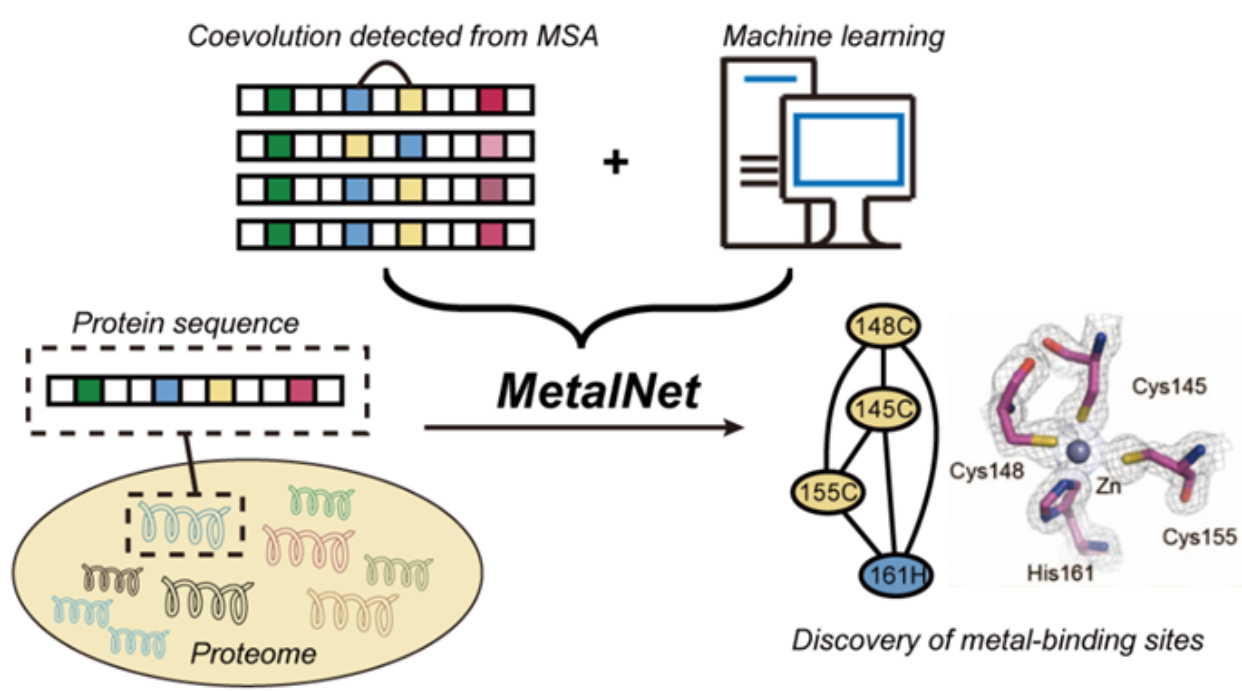
Fig. 1 Scheme of MetalNet.
The authors named the method MetalNet and applied it to predict metalloproteins and metal-binding sites in several proteomes. They first applied MetalNet to proteomes of four representative prokaryotic species and predicted 4,849 potential metalloproteins which significantly expands the currently annotated metalloproteomes. About half of these predicted proteins can be supported by structures in homologs, or can be indirectly supported from other databases (Fig. 2).
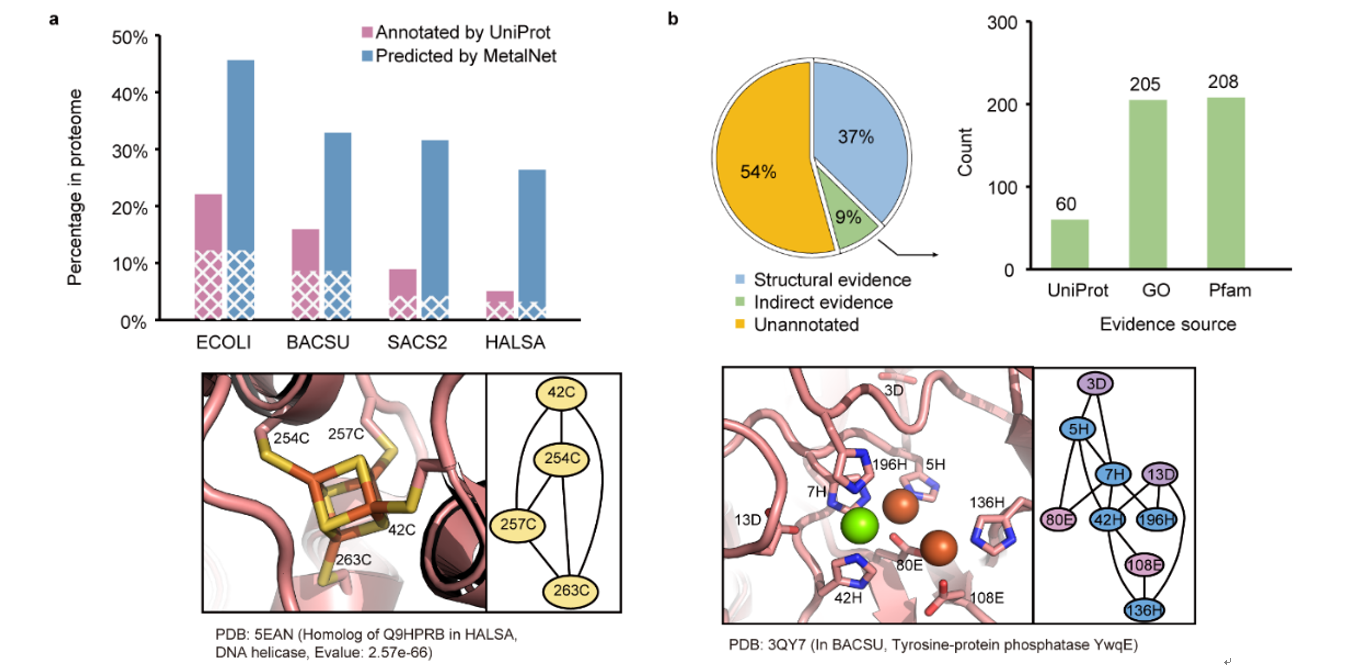
Fig. 2 Predictions of metalloproteins from representative prokaryotic species
They also biochemically and structurally validated previously unannotated metal-binding sites in several proteins, including apo-citrate lyase phosphoribosyl-dephospho-CoA transferase citX, an E.coli enzyme lacking structural or sequence homology to any known metalloprotein (PDB ID: 7DCM and 7DCN) (Fig. 3).
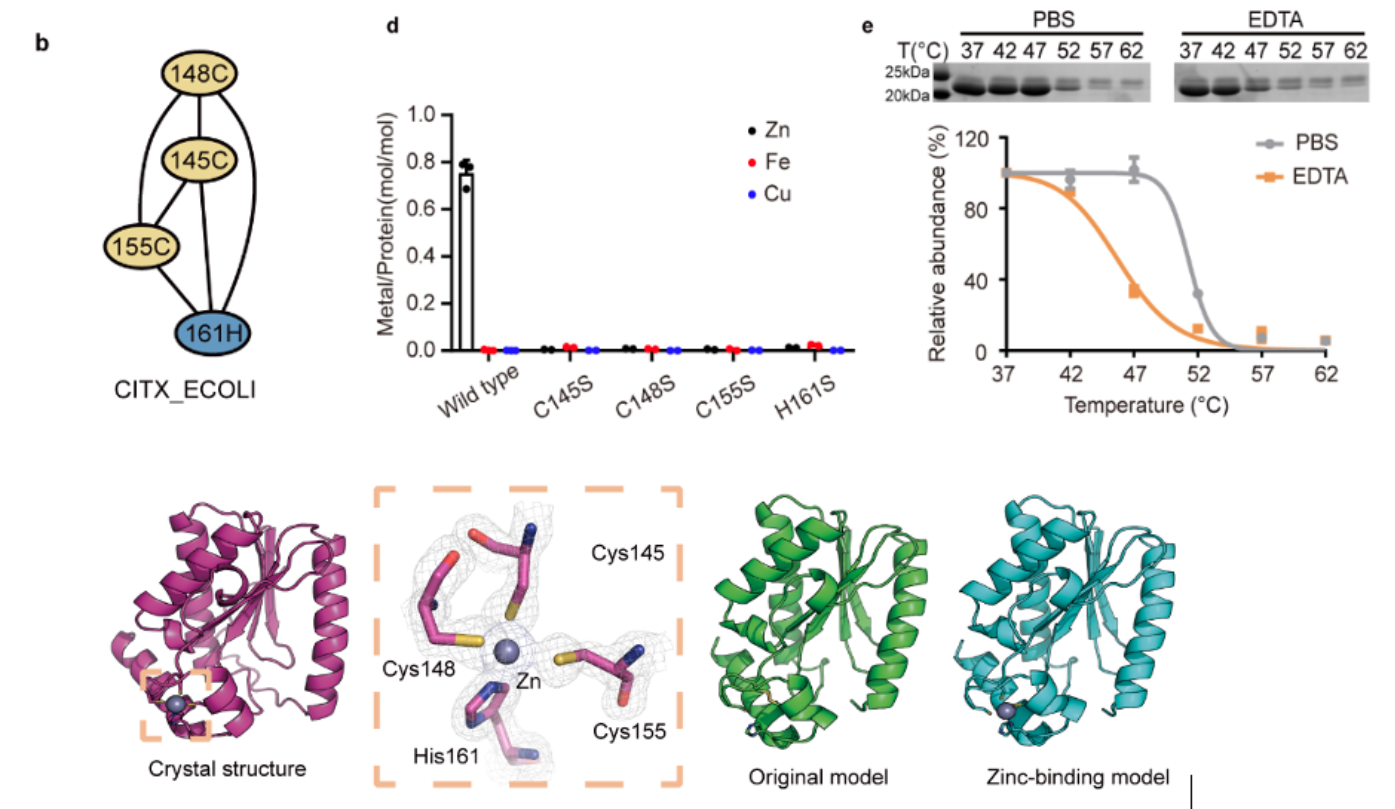
Fig. 3 Biochemical and structural validation of Citx as a zinc-binding protein
Finally, the authors applied MetalNet to the human spliceosome and found that all zinc-binding sites that were observed in the experimental structures could be recapitulated by MetalNet.
Prof. Xiao-Dong Su at State Key Laboratory of Protein and Plant Gene Research, and Biomedical Pioneering Innovation Center (BIOPIC), Peking University, Dr. Yuan Liu and Prof. Chu Wang at Synthetic and Functional Biomolecules Center, Beijing National Laboratory for Molecular Sciences, Key Laboratory of Bioorganic Chemistry and Molecular Engineering of Ministry of Education, Peking University are the co-corresponding authors of this paper. Dr. Yao Cheng, Dr. Haobo Wang, Dr. Yuan Liu from Prof. Chu Wang's research group at College of Chemistry and Molecular Engineering, and Dr. Hua Xu from the School of Life Sciences, Peking University, are the co-first authors of this paper. Collaborators including Bin Ma, Dr. Xuemin Chen, Dr. Xin Zeng and Xianghe Wang contributed to this project. This work was supported by the National Natural Science Foundation of China and Beijing National Laboratory for Molecular Sciences.
Original link for the paper: https://www.nature.com/articles/s41589-022-01223-z.